Sequential Selection¶
Theory¶
Sequential selection actually refers to an entire class of data-based methods. While there are many forms, we presently provide an implementation of the simplest two, sequential forward selection and sequential backward selection. A synopsis of these two methods, as well as several generalizations, can be found in Chapter 9 of Webb (2003). Sequential selection methods determine which predictors are important by evaluating model performance on a dataset where only some of the predictors are present. Predictors which, when present, improve the performance are typically considered important and predictors which, when removed, do not or only slightly degrade the performance are typically considered unimportant. In contrast with permutation importance, sequential selection methods train a new model at every step and are generally much more computationally expensive.
Sequential Forward Selection¶
Sequential forward selection iteratively adds predictors to the set of important predictors by taking the predictor at each step which most improves the performance of the model when added to the set of training predictors. This effectively determines the best 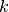 predictors for training a
predictors for training a 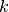 -predictor model. The process is demonstrated in Fig. 1: Sequential forward selection.
-predictor model. The process is demonstrated in Fig. 1: Sequential forward selection.
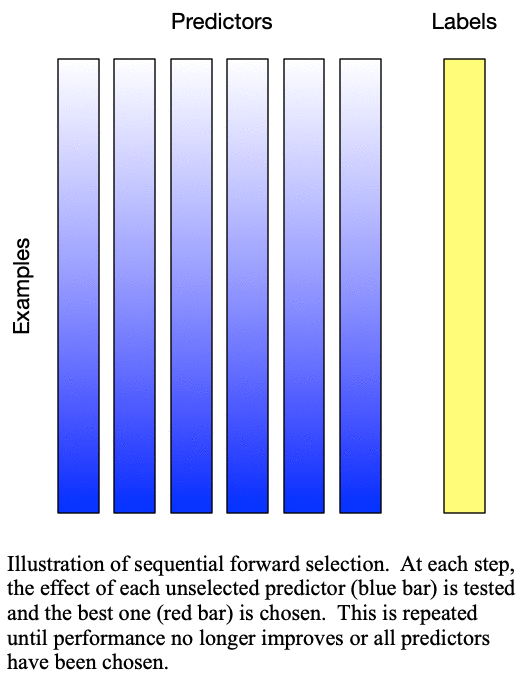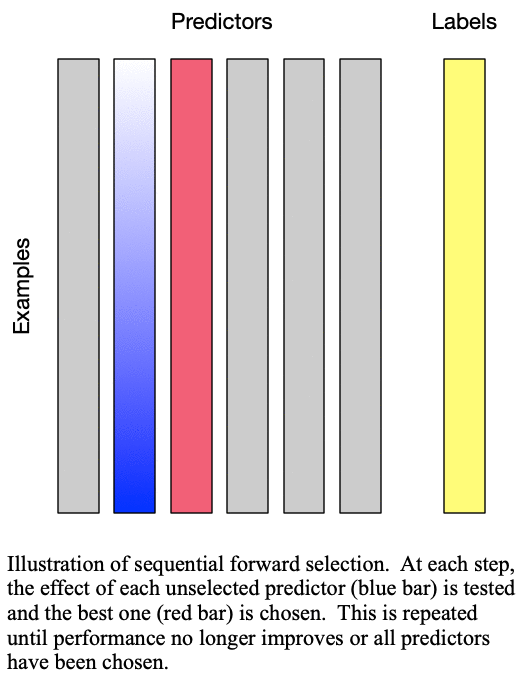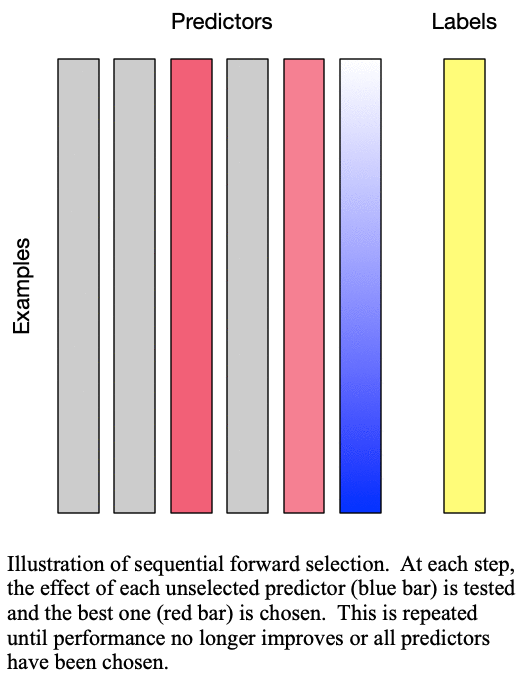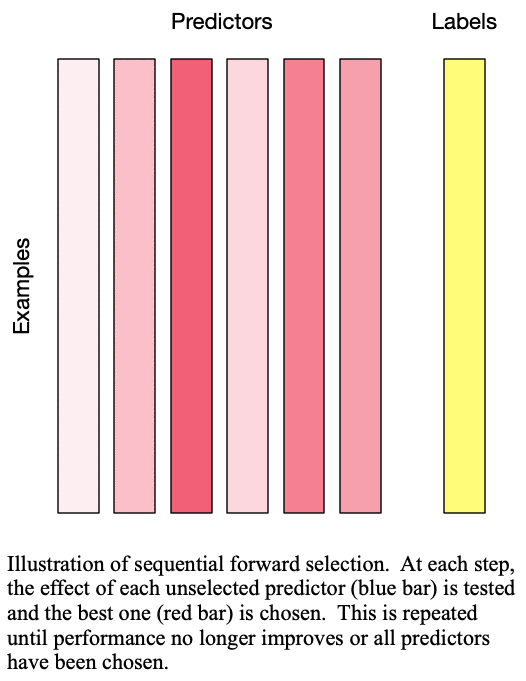
Fig. 1: Sequential forward selection adds the next-best predictor at each step
Sequential Backward Selection¶
Sequential backward selection iteratively removes variables from the set of important variables by taking the predictor at each step which least degrades the performance of the model when removed from the set of training predictors. This effectively determines the 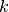 least important predictors. The process is demonstrated in Fig. 2: Sequential backward selection. A word of caution: sequential backward selection can take many times longer than sequential forward selection because it is training many more models with nearly complete sets of predictors. When there are more than 50 predictors, sequential backward selection often becomes computationally infeasible for some models.
least important predictors. The process is demonstrated in Fig. 2: Sequential backward selection. A word of caution: sequential backward selection can take many times longer than sequential forward selection because it is training many more models with nearly complete sets of predictors. When there are more than 50 predictors, sequential backward selection often becomes computationally infeasible for some models.
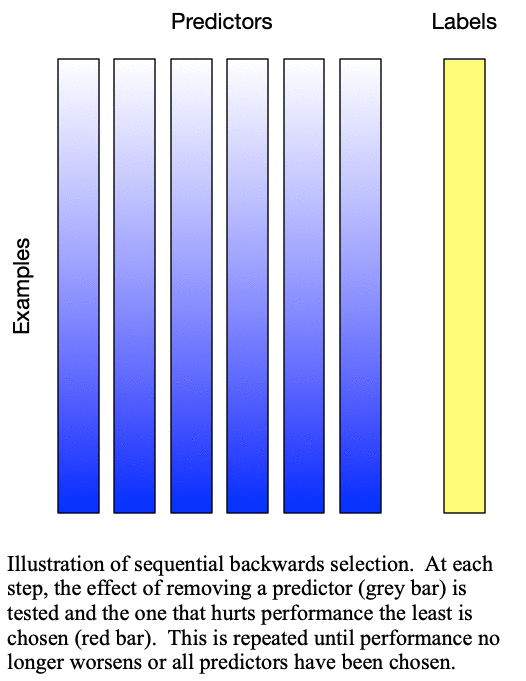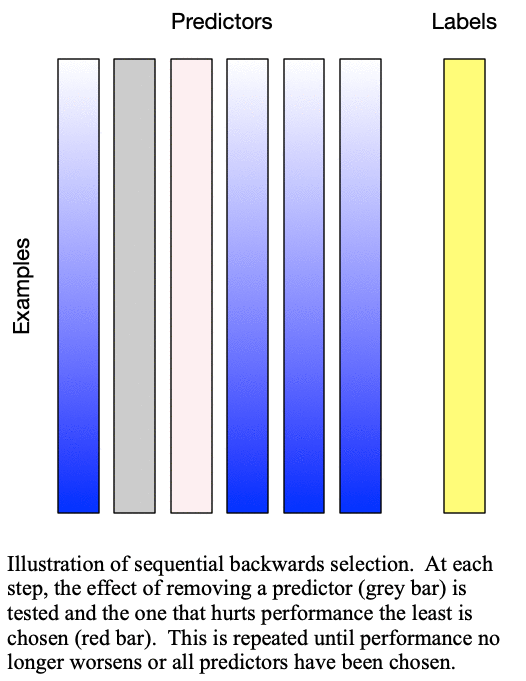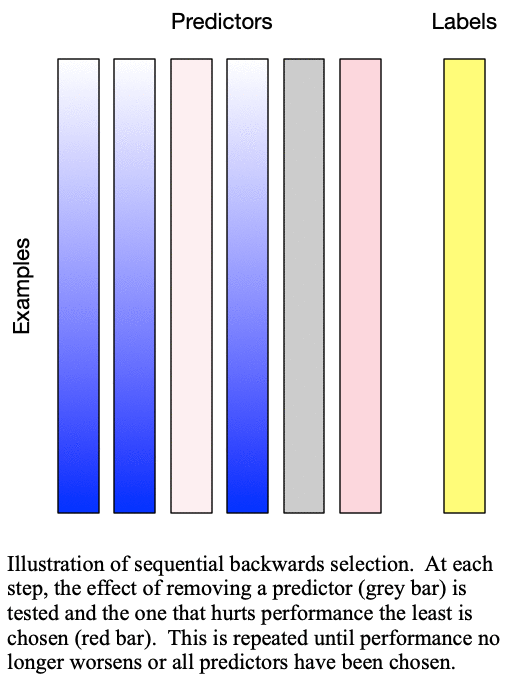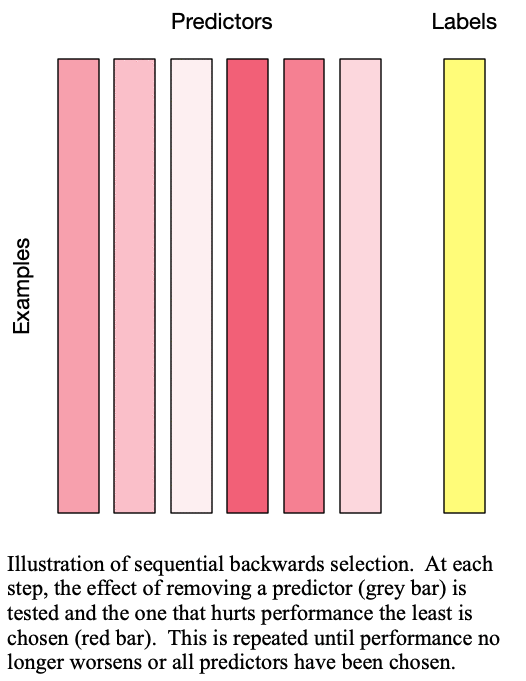
Fig. 2: Sequential backward selection removes the next-worst predictor at each step
Usage¶
As with all methods, we provide all sequential forward selection methods at two different levels of abstraction. For more information on the levels of abstraction and when to use each, please see Levels of Abstraction
Typically, when using a performance metric or skill score with any sequential
selection method, the scoring_strategy should be to maximize the performance.
On the other hand, when using an error or loss function, the scoring_strategy
should be to minimize the error or loss function.
Model-Based¶
-
PermutationImportance.sequential_selection.sklearn_sequential_forward_selection(model, training_data, scoring_data, evaluation_fn, scoring_strategy, variable_names=None, nimportant_vars=None, njobs=1, nbootstrap=None, subsample=1, **kwargs)[source] Performs sequential forward selection for a particular model,
scoring_data,evaluation_fn, and strategy for determining optimal variablesParameters: - model – a sklearn model
- training_data – a 2-tuple
(inputs, outputs)for training in thescoring_fn - scoring_data – a 2-tuple
(inputs, outputs)for scoring in thescoring_fn - evaluation_fn – a function which takes the deterministic or
probabilistic model predictions and scores them against the true
values. Must be of the form
(truths, predictions) -> some_valueProbably one of the metrics inPermutationImportance.metricsor sklearn.metrics - scoring_strategy – a function to be used for determining optimal
variables. Should be of the form
([some_value]) -> index - variable_names – an optional list for variable names. If not given, will use names of columns of data (if pandas dataframe) or column indices
- nimportant_vars – number of variables to compute importance for. Defaults to all variables
- njobs – an integer for the number of threads to use. If negative, will
use
num_cpus + njobs. Defaults to 1 - nbootstrap – number of times to perform scoring on each variable. Results over different bootstrap iterations are averaged. Defaults to 1
- subsample – number of elements to sample (with replacement) per bootstrap round. If between 0 and 1, treated as a fraction of the number of total number of events (e.g. 0.5 means half the number of events). If not specified, subsampling will not be used and the entire data will be used (without replacement)
- kwargs – all other kwargs will be passed on to the
evaluation_fn
Returns: PermutationImportance.result.ImportanceResultobject which contains the results for each run
Simple SFS Example¶
from sklearn.datasets import load_iris
from sklearn.metrics import accuracy_score
from sklearn.neural_network import MLPClassifier
from PermutationImportance import sklearn_sequential_forward_selection
# Separate out the last 20% for scoring data
iris = load_iris(return_X_y=False)
inputs = iris.get('data')
outputs = iris.get('target')
predictor_names = iris.get('feature_names')
training_inputs = inputs[:int(0.8 * len(inputs))]
training_outputs = outputs[:int(0.8 * len(outputs))]
scoring_inputs = inputs[int(0.8 * len(inputs)):]
scoring_outputs = outputs[int(0.8 * len(outputs)):]
# Train a quick neural net on the data
model = MLPClassifier(solver='lbfgs')
model.fit(training_inputs, training_outputs)
# Package the data into the right shape
training_data = (training_inputs, training_outputs)
scoring_data = (scoring_inputs, scoring_outputs)
# Use the sklearn_sequential_forward_selection to compute importances
result = sklearn_sequential_forward_selection(
model, training_data, scoring_data, accuracy_score, 'max',
variable_names=predictor_names)
# Get the Breiman-like singlepass results
print("Singlepass")
singlepass = result.retrieve_singlepass()
for predictor in singlepass.keys():
rank, score = singlepass[predictor]
print("Predictor: %s, Rank: %i, Score: %f" % (predictor, rank, score))
# Get the Lakshmanan-like multipass results
print("Multipass")
multipass = result.retrieve_multipass()
for predictor in multipass.keys():
rank, score = multipass[predictor]
print("Predictor: %s, Rank: %i, Score: %f" % (predictor, rank, score))
# Iterate over the (context, result) pairs
for i, (cntxt, res) in enumerate(result):
print("Context %i: %r" % (i, cntxt))
print("Result %i: %r" % (i, res))
Complex SFS Example¶
from sklearn.datasets import load_breast_cancer
from sklearn.ensemble import RandomForestClassifier
from PermutationImportance import sklearn_sequential_forward_selection
from PermutationImportance.metrics import peirce_skill_score
# Separate out the last 20% for scoring data
breast_cancer = load_breast_cancer(return_X_y=False)
inputs = breast_cancer.get('data')
outputs = breast_cancer.get('target')
predictor_names = breast_cancer.get('feature_names')
training_inputs = inputs[:int(0.8 * len(inputs))]
training_outputs = outputs[:int(0.8 * len(outputs))]
scoring_inputs = inputs[int(0.8 * len(inputs)):]
scoring_outputs = outputs[int(0.8 * len(outputs)):]
# Train a quick forest on the data
model = RandomForestClassifier(n_estimators=100, max_depth=4)
model.fit(training_inputs, training_outputs)
# Package the data into the right shape
training_data = (training_inputs, training_outputs)
scoring_data = (scoring_inputs, scoring_outputs)
# ----------- Version to use when only wanting singlepass results --------------
# Use the sklearn_sequential_forward_selection to compute importances
result = sklearn_sequential_forward_selection(
# argmax_of_mean handles bootstrapped metrics
model, training_data, scoring_data, peirce_skill_score, 'argmax_of_mean',
variable_names=predictor_names,
# sample (with replacement) 1*(number of samples) 5 times to compute metric distribution
# nbootstrap should typically be 1000, but this is kept small here for printing purposes
nbootstrap=5, subsample=1,
# only perform for the very top predictor (effectively means only compute singlepass results)
nimportant_vars=1)
# Get the Breiman-like singlepass results
print("Singlepass")
singlepass = result.retrieve_singlepass()
for predictor in singlepass.keys():
rank, score = singlepass[predictor]
print("Predictor: %s, Rank: %i, Score: %r" % (predictor, rank, score))
# Get the Lakshmanan-like multipass results
print("Multipass. This should only have 1 item and be not very useful")
multipass = result.retrieve_multipass()
for predictor in multipass.keys():
rank, score = multipass[predictor]
print("Predictor: %s, Rank: %i, Score: %r" % (predictor, rank, score))
# Iterate over the (context, result) pairs
for i, (cntxt, res) in enumerate(result):
print("Context %i: %r" % (i, cntxt))
print("Result %i: %r" % (i, res))
# ------------------------------------------------------------------------------
# ----------- Version to use when wanting multipass results --------------------
# Use the sklearn_sequential_forward_selection to compute importances
result = sklearn_sequential_forward_selection(
# argmax_of_mean handles bootstrapped metrics
model, training_data, scoring_data, peirce_skill_score, 'argmax_of_mean',
variable_names=predictor_names,
# sample (with replacement) 1*(number of samples) 5 times to compute metric distribution
# nbootstrap should typically be 1000, but this is kept small here for printing purposes
nbootstrap=5, subsample=1,
nimportant_vars=8) # only perform for the top 8 predictors
# Get the Breiman-like singlepass results
print("Singlepass")
singlepass = result.retrieve_singlepass()
for predictor in singlepass.keys():
rank, score = singlepass[predictor]
print("Predictor: %s, Rank: %i, Score: %r" % (predictor, rank, score))
# Get the Lakshmanan-like multipass results
print("Multipass. This should have exactly 8 items")
multipass = result.retrieve_multipass()
for predictor in multipass.keys():
rank, score = multipass[predictor]
print("Predictor: %s, Rank: %i, Score: %r" % (predictor, rank, score))
# Iterate over the (context, result) pairs
for i, (cntxt, res) in enumerate(result):
print("Context %i: %r" % (i, cntxt))
print("Result %i: %r" % (i, res))
# ------------------------------------------------------------------------------
# Use the plotting code in examples/plotting.py, found here:
# https://github.com/gelijergensen/PermutationImportance
try:
from plotting import plot_variable_importance
except Exception as e:
print("An error occurred while plotting. You probably don't have matplotlib installed")
print(e)
else:
plot_variable_importance(
result, 'example_sequential_forward_selection.png')
-
PermutationImportance.sequential_selection.sklearn_sequential_backward_selection(model, training_data, scoring_data, evaluation_fn, scoring_strategy, variable_names=None, nimportant_vars=None, njobs=1, nbootstrap=None, subsample=1, **kwargs)[source] Performs sequential backward selection for a particular model,
scoring_data,evaluation_fn, and strategy for determining optimal variablesParameters: - model – a sklearn model
- training_data – a 2-tuple
(inputs, outputs)for training in thescoring_fn - scoring_data – a 2-tuple
(inputs, outputs)for scoring in thescoring_fn - evaluation_fn –
a function which takes the deterministic or probabilistic model predictions and scores them against the true values. Must be of the form
(truths, predictions) -> some_valueProbably one of the metrics inPermutationImportance.metricsor sklearn.metrics - scoring_strategy – a function to be used for determining optimal
variables. Should be of the form
([some_value]) -> index - variable_names – an optional list for variable names. If not given, will use names of columns of data (if pandas dataframe) or column indices
- nimportant_vars – number of variables to compute importance for. Defaults to all variables
- njobs – an integer for the number of threads to use. If negative, will
use
num_cpus + njobs. Defaults to 1 - nbootstrap – number of times to perform scoring on each variable. Results over different bootstrap iterations are averaged. Defaults to 1
- subsample – number of elements to sample (with replacement) per bootstrap round. If between 0 and 1, treated as a fraction of the number of total number of events (e.g. 0.5 means half the number of events). If not specified, subsampling will not be used and the entire data will be used (without replacement)
- kwargs – all other kwargs will be passed on to the
evaluation_fn
Returns: PermutationImportance.result.ImportanceResultobject which contains the results for each run
Simple SBS Example¶
from sklearn.datasets import load_iris
from sklearn.metrics import accuracy_score
from sklearn.neural_network import MLPClassifier
from PermutationImportance import sklearn_sequential_backward_selection
# Separate out the last 20% for scoring data
iris = load_iris(return_X_y=False)
inputs = iris.get('data')
outputs = iris.get('target')
predictor_names = iris.get('feature_names')
training_inputs = inputs[:int(0.8 * len(inputs))]
training_outputs = outputs[:int(0.8 * len(outputs))]
scoring_inputs = inputs[int(0.8 * len(inputs)):]
scoring_outputs = outputs[int(0.8 * len(outputs)):]
# Train a quick neural net on the data
model = MLPClassifier(solver='lbfgs')
model.fit(training_inputs, training_outputs)
# Package the data into the right shape
training_data = (training_inputs, training_outputs)
scoring_data = (scoring_inputs, scoring_outputs)
# Use the sklearn_sequential_backward_selection to compute importances
result = sklearn_sequential_backward_selection(
model, training_data, scoring_data, accuracy_score, 'max',
variable_names=predictor_names)
# Get the Breiman-like singlepass results
print("Singlepass")
singlepass = result.retrieve_singlepass()
for predictor in singlepass.keys():
rank, score = singlepass[predictor]
print("Predictor: %s, Rank: %i, Score: %f" % (predictor, rank, score))
# Get the Lakshmanan-like multipass results
print("Multipass")
multipass = result.retrieve_multipass()
for predictor in multipass.keys():
rank, score = multipass[predictor]
print("Predictor: %s, Rank: %i, Score: %f" % (predictor, rank, score))
# Iterate over the (context, result) pairs
for i, (cntxt, res) in enumerate(result):
print("Context %i: %r" % (i, cntxt))
print("Result %i: %r" % (i, res))
Complex SBS Example¶
from sklearn.datasets import load_breast_cancer
from sklearn.ensemble import RandomForestClassifier
from PermutationImportance import sklearn_sequential_backward_selection
from PermutationImportance.metrics import peirce_skill_score
# Separate out the last 20% for scoring data
breast_cancer = load_breast_cancer(return_X_y=False)
inputs = breast_cancer.get('data')
outputs = breast_cancer.get('target')
predictor_names = breast_cancer.get('feature_names')
training_inputs = inputs[:int(0.8 * len(inputs))]
training_outputs = outputs[:int(0.8 * len(outputs))]
scoring_inputs = inputs[int(0.8 * len(inputs)):]
scoring_outputs = outputs[int(0.8 * len(outputs)):]
# Train a quick forest on the data
model = RandomForestClassifier(n_estimators=100, max_depth=4)
model.fit(training_inputs, training_outputs)
# Package the data into the right shape
training_data = (training_inputs, training_outputs)
scoring_data = (scoring_inputs, scoring_outputs)
# ----------- Version to use when only wanting singlepass results --------------
# Use the sklearn_sequential_backward_selection to compute importances
result = sklearn_sequential_backward_selection(
# argmax_of_mean handles bootstrapped metrics
model, training_data, scoring_data, peirce_skill_score, 'argmax_of_mean',
variable_names=predictor_names,
# sample (with replacement) 1*(number of samples) 5 times to compute metric distribution
# nbootstrap should typically be 1000, but this is kept small here for printing purposes
nbootstrap=5, subsample=1,
# only perform for the very top predictor (effectively means only compute singlepass results)
nimportant_vars=1)
# Get the Breiman-like singlepass results
print("Singlepass")
singlepass = result.retrieve_singlepass()
for predictor in singlepass.keys():
rank, score = singlepass[predictor]
print("Predictor: %s, Rank: %i, Score: %r" % (predictor, rank, score))
# Get the Lakshmanan-like multipass results
print("Multipass. This should only have 1 item and be not very useful")
multipass = result.retrieve_multipass()
for predictor in multipass.keys():
rank, score = multipass[predictor]
print("Predictor: %s, Rank: %i, Score: %r" % (predictor, rank, score))
# Iterate over the (context, result) pairs
for i, (cntxt, res) in enumerate(result):
print("Context %i: %r" % (i, cntxt))
print("Result %i: %r" % (i, res))
# ------------------------------------------------------------------------------
# ----------- Version to use when wanting multipass results --------------------
# Use the sklearn_sequential_backward_selection to compute importances
result = sklearn_sequential_backward_selection(
# argmax_of_mean handles bootstrapped metrics
model, training_data, scoring_data, peirce_skill_score, 'argmax_of_mean',
variable_names=predictor_names,
# sample (with replacement) 1*(number of samples) 5 times to compute metric distribution
# nbootstrap should typically be 1000, but this is kept small here for printing purposes
nbootstrap=5, subsample=1,
nimportant_vars=8) # only perform for the top 8 predictors
# Get the Breiman-like singlepass results
print("Singlepass")
singlepass = result.retrieve_singlepass()
for predictor in singlepass.keys():
rank, score = singlepass[predictor]
print("Predictor: %s, Rank: %i, Score: %r" % (predictor, rank, score))
# Get the Lakshmanan-like multipass results
print("Multipass. This should have exactly 8 items")
multipass = result.retrieve_multipass()
for predictor in multipass.keys():
rank, score = multipass[predictor]
print("Predictor: %s, Rank: %i, Score: %r" % (predictor, rank, score))
# Iterate over the (context, result) pairs
for i, (cntxt, res) in enumerate(result):
print("Context %i: %r" % (i, cntxt))
print("Result %i: %r" % (i, res))
# ------------------------------------------------------------------------------
# Use the plotting code in examples/plotting.py, found here:
# https://github.com/gelijergensen/PermutationImportance
try:
from plotting import plot_variable_importance
except Exception as e:
print("An error occurred while plotting. You probably don't have matplotlib installed")
print(e)
else:
plot_variable_importance(
result, 'example_sequential_backward_selection.png')
Method-Specific¶
-
PermutationImportance.sequential_selection.sequential_forward_selection(training_data, scoring_data, scoring_fn, scoring_strategy, variable_names=None, nimportant_vars=None, njobs=1)[source] Performs sequential forward selection over data given a particular set of functions for scoring and determining optimal variables
Parameters: - training_data – a 2-tuple
(inputs, outputs)for training in thescoring_fn - scoring_data – a 2-tuple
(inputs, outputs)for scoring in thescoring_fn - scoring_fn – a function to be used for scoring. Should be of the form
(training_data, scoring_data) -> some_value - scoring_strategy – a function to be used for determining optimal
variables. Should be of the form
([some_value]) -> index - variable_names – an optional list for variable names. If not given, will use names of columns of data (if pandas dataframe) or column indices
- nimportant_vars – number of variables to compute importance for. Defaults to all variables
- njobs – an integer for the number of threads to use. If negative, will
use
num_cpus + njobs. Defaults to 1
Returns: PermutationImportance.result.ImportanceResultobject which contains the results for each run- training_data – a 2-tuple
SFS Example¶
from sklearn.datasets import load_iris
from sklearn.metrics import accuracy_score
from sklearn.neural_network import MLPClassifier
from PermutationImportance import sequential_forward_selection
# Separate out the last 20% for scoring data
iris = load_iris(return_X_y=False)
inputs = iris.get('data')
outputs = iris.get('target')
predictor_names = iris.get('feature_names')
training_inputs = inputs[:int(0.8 * len(inputs))]
training_outputs = outputs[:int(0.8 * len(outputs))]
scoring_inputs = inputs[int(0.8 * len(inputs)):]
scoring_outputs = outputs[int(0.8 * len(outputs)):]
def score_model(training_data, scoring_data):
"""Custom function to use for scoring. Notice that because this is
sequential selection, we need to retrain the model each time
:param training_data: (training_inputs, training_outputs)
:param scoring_data: (scoring_inputs, scoring_outputs)
"""
training_ins, training_outs = training_data
scoring_ins, scoring_outs = scoring_data
# Some model we are interested in
model = MLPClassifier(solver='lbfgs')
model.fit(training_ins, training_outs)
return accuracy_score(scoring_outs, model.predict(scoring_ins))
# Package the data into the right shape
training_data = (training_inputs, training_outputs)
scoring_data = (scoring_inputs, scoring_outputs)
# Use the sequential_forward_selection to compute importances
result = sequential_forward_selection(
training_data, scoring_data, score_model, 'max',
variable_names=predictor_names)
# Get the Breiman-like singlepass results
print("Singlepass")
singlepass = result.retrieve_singlepass()
for predictor in singlepass.keys():
rank, score = singlepass[predictor]
print("Predictor: %s, Rank: %i, Score: %f" % (predictor, rank, score))
# Get the Lakshmanan-like multipass results
print("Multipass")
multipass = result.retrieve_multipass()
for predictor in multipass.keys():
rank, score = multipass[predictor]
print("Predictor: %s, Rank: %i, Score: %f" % (predictor, rank, score))
# Iterate over the (context, result) pairs
for i, (cntxt, res) in enumerate(result):
print("Context %i: %r" % (i, cntxt))
print("Result %i: %r" % (i, res))
-
PermutationImportance.sequential_selection.sequential_backward_selection(training_data, scoring_data, scoring_fn, scoring_strategy, variable_names=None, nimportant_vars=None, njobs=1)[source] Performs sequential backward selection over data given a particular set of functions for scoring and determining optimal variables
Parameters: - training_data – a 2-tuple
(inputs, outputs)for training in thescoring_fn - scoring_data – a 2-tuple
(inputs, outputs)for scoring in thescoring_fn - scoring_fn – a function to be used for scoring. Should be of the form
(training_data, scoring_data) -> some_value - scoring_strategy – a function to be used for determining optimal
variables. Should be of the form
([some_value]) -> index - variable_names – an optional list for variable names. If not given, will use names of columns of data (if pandas dataframe) or column indices
- nimportant_vars – number of variables to compute importance for. Defaults to all variables
- njobs – an integer for the number of threads to use. If negative, will
use
num_cpus + njobs. Defaults to 1
Returns: PermutationImportance.result.ImportanceResultobject which contains the results for each run- training_data – a 2-tuple
SBS Example¶
from sklearn.datasets import load_iris
from sklearn.metrics import accuracy_score
from sklearn.neural_network import MLPClassifier
from PermutationImportance import sequential_backward_selection
# Separate out the last 20% for scoring data
iris = load_iris(return_X_y=False)
inputs = iris.get('data')
outputs = iris.get('target')
predictor_names = iris.get('feature_names')
training_inputs = inputs[:int(0.8 * len(inputs))]
training_outputs = outputs[:int(0.8 * len(outputs))]
scoring_inputs = inputs[int(0.8 * len(inputs)):]
scoring_outputs = outputs[int(0.8 * len(outputs)):]
def score_model(training_data, scoring_data):
"""Custom function to use for scoring. Notice that because this is
sequential selection, we need to retrain the model each time
:param training_data: (training_inputs, training_outputs)
:param scoring_data: (scoring_inputs, scoring_outputs)
"""
training_ins, training_outs = training_data
scoring_ins, scoring_outs = scoring_data
# Some model we are interested in
model = MLPClassifier(solver='lbfgs')
model.fit(training_ins, training_outs)
return accuracy_score(scoring_outs, model.predict(scoring_ins))
# Package the data into the right shape
training_data = (training_inputs, training_outputs)
scoring_data = (scoring_inputs, scoring_outputs)
# Use the sequential_backward_selection to compute importances
result = sequential_backward_selection(
training_data, scoring_data, score_model, 'max',
variable_names=predictor_names)
# Get the Breiman-like singlepass results
print("Singlepass")
singlepass = result.retrieve_singlepass()
for predictor in singlepass.keys():
rank, score = singlepass[predictor]
print("Predictor: %s, Rank: %i, Score: %f" % (predictor, rank, score))
# Get the Lakshmanan-like multipass results
print("Multipass")
multipass = result.retrieve_multipass()
for predictor in multipass.keys():
rank, score = multipass[predictor]
print("Predictor: %s, Rank: %i, Score: %f" % (predictor, rank, score))
# Iterate over the (context, result) pairs
for i, (cntxt, res) in enumerate(result):
print("Context %i: %r" % (i, cntxt))
print("Result %i: %r" % (i, res))